This vignette focuses on the modern bipl5 workflow for
principal component analysis (PCA) biplots:
init_biplot(...) |> scale_mds(type = "pca", ...)The mathematical background is the same background documented in
wrap_bipl5.PCA(), but the examples below use the newer
specification-based workflow because it is the most natural entry point
for package users who want to build a single display first and then
extend it later.
The goals of this vignette are to:
- construct a PCA biplot with
init_biplot()andscale_mds() - show the printed structure of the resulting
bipl5_biplot - add extra
mdsDisplayswithappend_mdsDisplay() - demonstrate
format_samples()with both one and two categorical stratifications - explain the PCA fit measures stored in
fit_measures - preserve the current long-form documentation of the PCA biplot mathematics
Building a PCA biplot
init_biplot() stores the data and preprocessing choices,
while scale_mds() fits and compiles the requested biplot.
If the input data frame contains categorical columns, those columns are
retained inside the specification object so they can be used later by
format_samples().
iris2 <- iris
iris2$Band <- factor(
rep(c("class1", "class2", "class3", "class4"), length.out = nrow(iris2))
)
pca_spec <- init_biplot(iris2, scale = TRUE)
bp_pca <- pca_spec |>
scale_mds(
type = "pca",
eigenvectors = c(1, 2),
group_aes = iris2$Species,
show_group_means = TRUE
)Immediately after scale_mds(), the object contains one
compiled display and a PCA fit-measures branch:
bp_pca
#> bipl5_biplot [PCA]
#> ├── mdsDisplay_12 [PC 1 & 2] <bipl5_mdsDisplay>
#> │ ├── Data <bipl5_data>
#> │ │ ├── sample_coordinates [150 x 2]
#> │ │ ├── axes_coordinates [4 axes]
#> │ │ └── translated_axes_coordinates
#> │ ├── trace_data [35 traces]
#> │ └── annotations [140 items]
#> └── fit_measures <bipl5_fitmeasures>
#> ├── CumPred [5 traces]
#> ├── CumAd [4 traces]
#> ├── VarExp [5 traces]
#> ├── Scree [1 traces]
#> └── fit_table_12 [PC 1 & 2]The default PCA plot is interactive. The left-hand side is the biplot itself, while the right-hand side fit panel can be opened from the widget controls.
plot(bp_pca)Adding more mdsDisplays
The scale_mds() workflow deliberately compiles only the
requested pair of principal components. This keeps the initial object
light, and it mirrors the fact that a user often wants to start with
just one view. Additional views can then be added with
append_mdsDisplay().
Here we extend the original PC 1 & 2 display to also
include PC 1 & 3 and PC 2 & 3:
bp_pca_full <- bp_pca |>
append_mdsDisplay(c(1, 3)) |>
append_mdsDisplay(c(2, 3))The printed tree now reflects three stored displays and one fit table for each pair:
bp_pca_full
#> bipl5_biplot [PCA]
#> ├── mdsDisplay_12 [PC 1 & 2] <bipl5_mdsDisplay>
#> │ ├── Data <bipl5_data>
#> │ │ ├── sample_coordinates [150 x 2]
#> │ │ ├── axes_coordinates [4 axes]
#> │ │ └── translated_axes_coordinates
#> │ ├── trace_data [35 traces]
#> │ └── annotations [140 items]
#> ├── mdsDisplay_13 [PC 1 & 3] <bipl5_mdsDisplay>
#> │ ├── Data <bipl5_data>
#> │ │ ├── sample_coordinates [150 x 2]
#> │ │ ├── axes_coordinates [4 axes]
#> │ │ └── translated_axes_coordinates
#> │ ├── trace_data [35 traces]
#> │ └── annotations [150 items]
#> ├── mdsDisplay_23 [PC 2 & 3] <bipl5_mdsDisplay>
#> │ ├── Data <bipl5_data>
#> │ │ ├── sample_coordinates [150 x 2]
#> │ │ ├── axes_coordinates [4 axes]
#> │ │ └── translated_axes_coordinates
#> │ ├── trace_data [35 traces]
#> │ └── annotations [160 items]
#> └── fit_measures <bipl5_fitmeasures>
#> ├── CumPred [5 traces]
#> ├── CumAd [4 traces]
#> ├── VarExp [5 traces]
#> ├── Scree [1 traces]
#> ├── fit_table_12 [PC 1 & 2]
#> ├── fit_table_13 [PC 1 & 3]
#> └── fit_table_23 [PC 2 & 3]The plot now exposes a dropdown for switching between the stored PC pairs.
plot(bp_pca_full)Formatting samples
format_samples() does not refit the PCA model. It
rebuilds the sample-trace block inside each stored
mdsDisplay, so it is the right tool when the ordination is
fixed but the display grouping needs to change.
We begin with a colour stratification by species:
bp_pca_species <- bp_pca_full |>
format_samples(
stratify = "col",
by = Species,
col = c("tomato", "steelblue", "darkgreen")
)We then add a second categorical stratification through plotting
symbols. Since Band is different from Species,
the plotted observation traces are rebuilt internally as the observed
Species x Band combinations, while the legend is kept
readable by showing separate legend sections for the colour and symbol
variables.
bp_pca_dual <- bp_pca_species |>
format_samples(
stratify = "symbol",
by = Band,
pch = c(15, 16, 17, 18)
)The formatted object remains a bipl5_biplot, so it can
still be printed, subset, extended, and plotted as usual.
plot(bp_pca_dual)PCA fit measures
PCA biplots are the richest objects in the current package because
they carry a full fit_measures branch. The print method
makes the available fit objects explicit:
bp_pca_full$fit_measures
#> bipl5_fitmeasures
#> ├── CumPred [5 traces]
#> ├── CumAd [4 traces]
#> ├── VarExp [5 traces]
#> ├── Scree [1 traces]
#> ├── fit_table_12 [PC 1 & 2]
#> ├── fit_table_13 [PC 1 & 3]
#> └── fit_table_23 [PC 2 & 3]In the interactive PCA widget, these are exposed through the “Measures of Fit” button and the fit-panel dropdown. The fit objects are:
-
CumPred: cumulative predictivity across increasing subspace dimension. Each line tracks one variable’s cumulative axis predictivity, and the weighted mean line equals the overall display quality. -
CumAd: cumulative adequacy across increasing subspace dimension. This shows how well each variable direction is represented by the accumulated loading structure, irrespective of the final two-dimensional display pair. -
VarExp: variance explained. In the current implementation this combines a line for the individual proportion of variance explained by each principal component with stacked variable-level contributions. -
Scree: the eigenvalue profile across the principal components. -
fit_table_ij: the pair-specific summary table for the active display. Each table reports variable-level adequacy and predictivity for that specificPC i & jmap.
If you want one of the graph-based fit objects on its own, you can extract it and plot it directly:
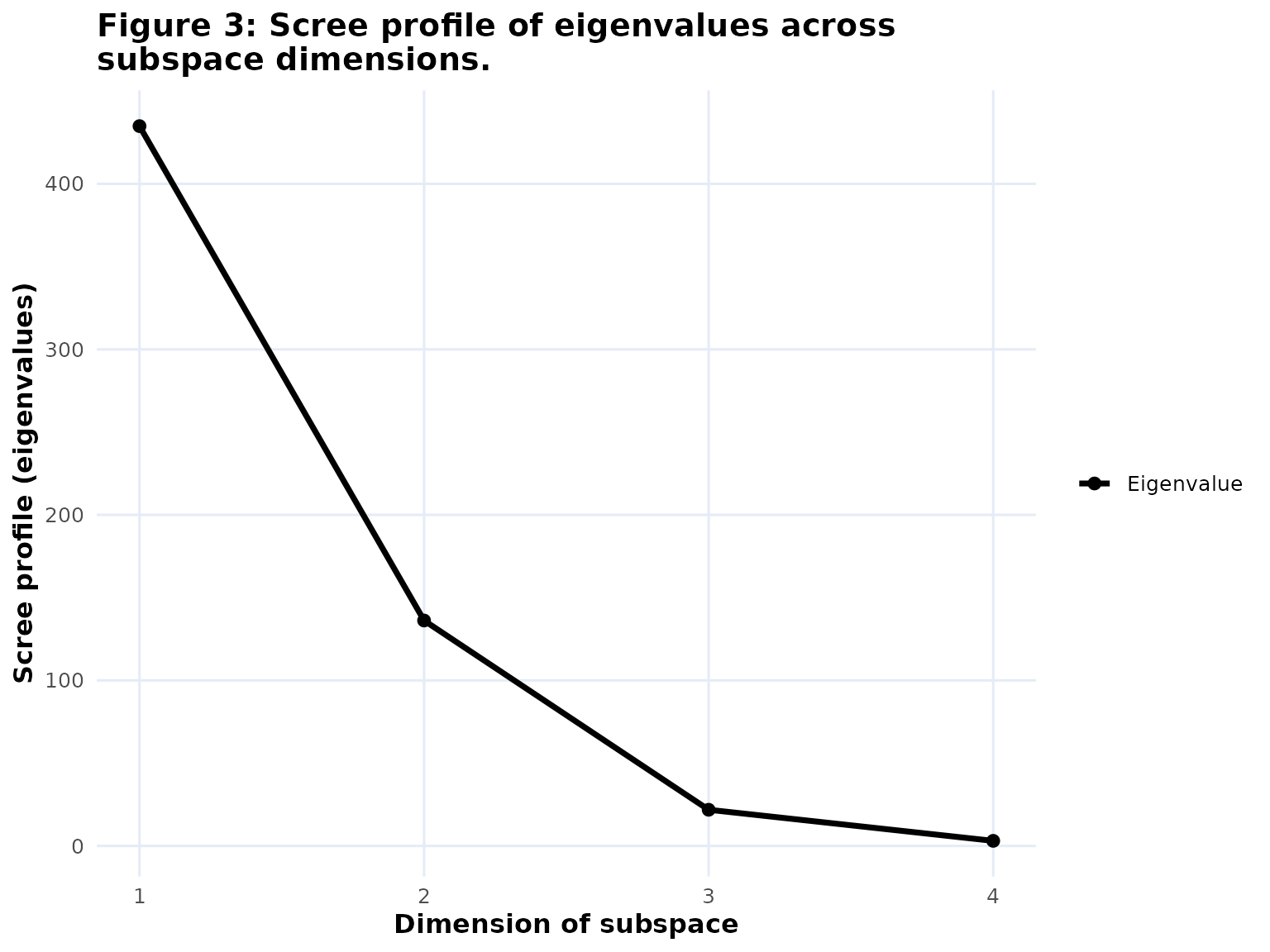
The summary table is not a separate bipl5_fit graph
object, because it is already stored in the fit_measures
branch as a plotly table trace. It updates when the active
mdsDisplay changes.
Mathematical background
This section preserves the current long-form PCA documentation in narrative form.
Let denote the processed data matrix stored in the input object after centring and any optional scaling. PCA is performed on that processed matrix, so the singular value decomposition is
where , , and with and . The principal component score vectors are
Suppose the displayed pair is with . Let be the diagonal selector matrix with ones in positions and . Then the two-dimensional fitted matrix is
Equivalently, with , , and containing only the selected components,
For the ordinary PCA biplot the displayed sample coordinates are
so that
If is the displayed direction for variable , then the fitted value for observation on variable is
and the calibrated axis point corresponding to marker value is
For a correlation biplot the factorization can instead be written as
so the displayed sample coordinates are and the displayed variable directions are the rows of . In that setting the variable directions are tuned to variable-correlation structure rather than raw score geometry.
Two orthogonality relations are central:
and
where . The first justifies sample-side fit measures (Type A orthogonality), and the second justifies variable-side fit measures (Type B orthogonality).
For sample , the sample predictivity is
For variable , the axis predictivity is
The overall quality of the displayed PCA plane is
Because of orthogonality, this same quantity can be read as a weighted average of either the sample predictivities or the axis predictivities. On the variable side,
When the processed variables all have equal sums of squares, the overall quality is simply the average axis predictivity. This is one reason standardized PCA is often easy to interpret.
PCA also gives a clean per-component decomposition. Since
the two displayed rank-1 pieces are orthogonal, and therefore
Likewise,
so both axis-side and sample-side quality can be broken down into the two displayed components.
The package documentation also distinguishes these sum-of-squares measures from the direct-reading diagnostics of Alves (2012). If is the displayed direction of variable , then the read-off value on that axis for sample is
The direct-reading error is
where is the standard deviation used in preprocessing, and the mean axis-level direct-reading error is
These quantities answer a different question from predictivity. Predictivity asks how much sum of squares is reproduced by the displayed plane. The Alves diagnostics ask how accurately values can be read directly from the plotted calibrated axes.
References
Alves, M. R. (2012). Evaluation of the predictive power of biplot axes to automate the construction and layout of biplots based on the accuracy of direct readings from common outputs of multivariate analyses: application to principal component analysis. Journal of Chemometrics, 26(5), 180-190.
Eckart, C. and Young, G. (1936). The approximation of one matrix by another of lower rank. Psychometrika, 1, 211-218.
Gabriel, K. R. (1971). The biplot graphical display of matrices with application to principal component analysis. Biometrika, 58(3), 453-467.
Gardner-Lubbe, S., le Roux, N. J. and Gower, J. C. (2008). Measures of fit in principal component and canonical variate analyses. Journal of Applied Statistics, 35(9), 947-965.
Gower, J. C. and Hand, D. J. (1996). Biplots. London: Chapman and Hall.
Gower, J. C., Lubbe, S. and le Roux, N. J. (2011). Understanding Biplots. Chichester: Wiley.
Greenacre, M. (2010). Biplots in Practice. Bilbao: BBVA Foundation.